7 Free Image Analysis Software Tools for Microscopy
Originally published by StressMarq Biosciences Jan 2015 | Updated July 2025
Microscopy image analysis is a critical step in biological research. Fortunately, there are several free tools available that offer powerful features for researchers. Here’s an updated list of 7 free image analysis software tools for microscopy, including two new additions from the 2020s.
✅ Original Tools (Reviewed)
1. ImageJ
Still highly relevant. Remains a go-to platform for image analysis in life sciences.
2. CellProfiler
Still maintained and widely used.
3. NeuronStudio
Outdated. Last updated in 2009 and requires legacy Windows OS. Consider replacing or noting its limitations.
4. Vaa3D
Still active, especially for 3D visualization and neuron tracing. Vaa3D (Visualization-Assisted Analysis 3D) is an open-source software platform designed for the visualization and analysis of large-scale, multidimensional biological image data. It excels in handling 3D, 4D, and even higher-dimensional datasets, making it especially useful for applications like neuron tracing, brain mapping, and volumetric microscopy.
5. Icy
Still supported and offers a user-friendly interface for bioimage analysis. Icy is an open-source bioimage analysis platform designed to be both powerful and user-friendly. Developed by the BioImage Analysis unit at Institut Pasteur, Icy supports a wide range of image processing tasks including segmentation, tracking, and 3D visualization.
🆕 New Tools to Add (2020s)
6. CellDetective
A modern, AI-enhanced tool for segmentation, tracking, and analysis of time-lapse microscopy datasets. Designed for immunology and cell biology, it’s open-source and user-friendly.
7. LABKIT
A powerful Fiji plugin for big image data. Offers manual and automated segmentation, supports 2D/3D/timelapse, and is optimized for large datasets.
These tools provide a wide range of capabilities for researchers working with microscopy data. Whether you’re analyzing cells, neurons, or tissues, there’s a free tool to help you get the job done.
Original blog (Jan 2015)
If your lab was like mine was, you struggle to afford new antibodies, let alone expensive image analysis software programs like Neurolucida to analyze your data.
Out of sheer necessity to analyze a massive image set before I turned grey, I scrounged around for free open source software programs to help analyze my confocal microscopy image stacks.
It took me years (actually!) to find some of them, so I thought I would pay-it-forward by sharing them with you! Kudos to everyone who takes the time to create these free open source programs.
Here are my Top 5 favorite time/life savers:
1. ImageJ/FIJI – The Image Multi-Tool
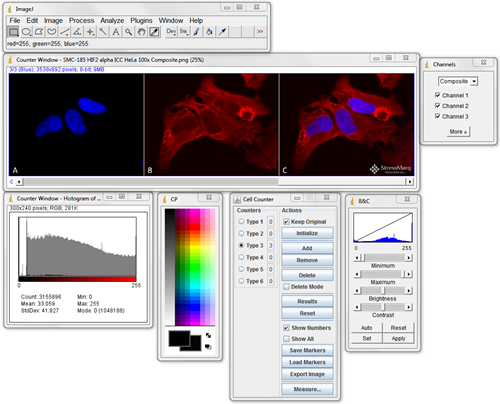
ImageJ should be the first program you become familiar with when looking for image analysis software. Download it, search through the plugins to see what’s available and test them out. Being proficient at using ImageJ is essential for most image processing and analysis.
It can do simple things like crop, label, and alter the brightness and contrast of fluorescence images. It can also easily handle 3D stacks of confocal microscopy images, and perform complex quantitative analysis.
For more information visit their website: http://imagej.nih.gov/ij/
FIJI (FIJI Is Just ImageJ) is a bundle of the top plugins available for ImageJ. Downloading FIJI instead of ImageJ just starts you out with more base plugins so you don’t have to hunt around and add them to your ImageJ program. There were a few plugins that were only available in FIJI. The select grouping of plugins is geared for neuroscience.
For more information visit their website: http://fiji.sc/Fiji
2. Cell Profiler/Cell Analyst – Image Batch Processing and Data Analysis
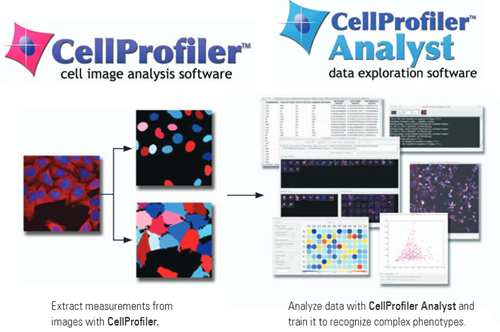
Cell Profiler can extract quantitative measurements from thousands of images through a custom pipeline that can first process and then analyze your images.
It can quantify antibody fluorescence intensity, colocalization, cell density, or even track cells through time-lapse movies. It’s as simple as setting up your pipeline and then going for a coffee.
I’ve never used Cell Analyst but I’ve heard good things about its ability to handle large datasets. Give it a try and let me know what you think.
For more information visit their website: http://www.cellprofiler.org/
3. Neuronstudio – Automatic Neuron Tracing Super-Tool
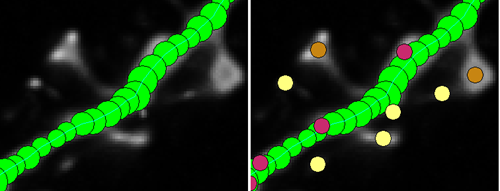
If you need to trace the dendritic arbor of a neuron then this is by far the best free open source program that I came across. Don’t be thrown by the lack of updates since 2009 or the fact that you need Windows 2000/XP/Vista to run it. It has a comprehensive help manual that will walk you through everything you need to know.
It has manual, semi-manual, and automatic tracing modes for complete 3D reconstruction. All-around the program is a breeze to use, and does a fantastic job of automatically tracing through a stack of confocal images based on the fluorescence intensity.
You can even find errors in your 3D reconstruction by flipping the trace file and image stack into a different dimension to look for anomalies in the trace, and then easily fix them by dragging the point to the correct location.
It will also perform a quantitative analysis on dendritic spine morphology. It can distinguish between different types, or you can set your own classification parameters.
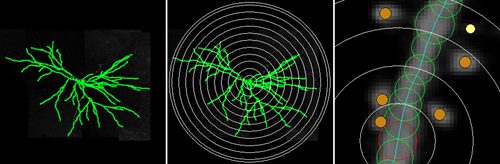
It can perform a basic Sholl Analysis on the dendritic arbor and output branch order data, including spine counts.
The program also lets you output your trace as a .swc file, which allows you to reconstruct or analyze the traced neuron in other programs, such as L-measure (below).
For more information visit their website: http://research.mssm.edu/cnic/tools-ns.html
4. Volume Integration and Alignment System (VIAS) – Image Stack Alignment Software
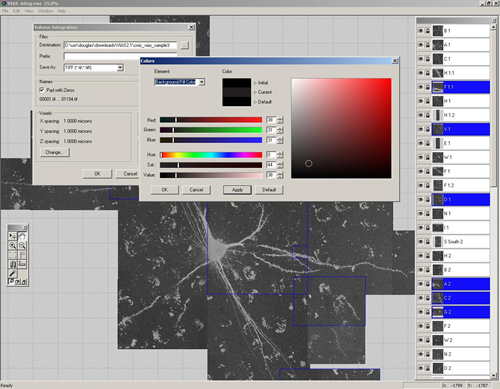
VIAS enables you to tile multiple confocal microscopy image stacks into a single 3D image dataset. This is a software program made by the same group that created Neuronstudio. It has a similar intuitive user interface and a step-by-step online guide.
It allows for real-time interaction with the stacks, therefore you can easily drag the stacks around until they are perfectly aligned. An auto-alignment feature helps to tweak the alignment for a seamless result.
If your image stacks are of different depths, they can be easily aligned in the z-dimension. Once finished you can export as a .tif to use the new stack in Neuronstudio, or ImageJ.
For more information visit their website: http://research.mssm.edu/cnic/tools-vias.html. Sorry, I cannot locate the current link to this software. Please comment if you know where it can be downloaded.
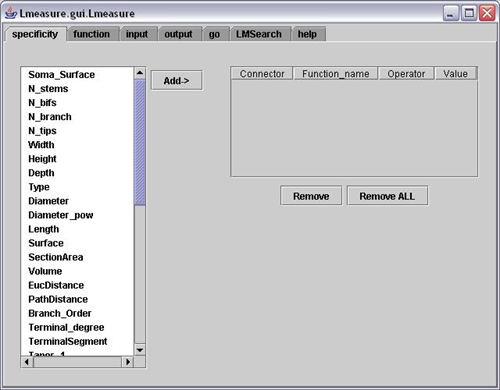
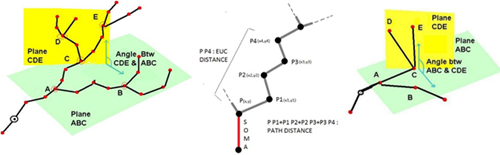
5. L-measure – Quantitative Morphological Measurements to the Max
L-measure was my tool of choice to extract more complex quantitative measurements from my neuronal reconstructions. It has a huge range of morphological measurements that you cannot find built into other programs.
It will take your morphological analysis to a whole new level by analyzing the volume of dendrites, the asymmetry of branches or even the angle between branches.
The online help guide is fantastic and goes into great depth to explain the complex measurements. The user interface is super simple and allows for multiple traces files (.swc format) to be analyzed at once and will output one file for your subsequent statistical analysis.
For more information visit their online help guide: http://cng.gmu.edu:8080/Lm/help/index.htm
To Download L-measure visit their website: http://cng.gmu.edu:8080/Lm/
This is just a list of the programs that I loved during my graduate work. There are plenty of other great free open source image analysis programs available.
Have questions about any of these programs? Ask them in the comments section below!



Digital microscopes are truly fascinating tools! The ability to explore the microscopic world with high-resolution imaging and digital capabilities opens up endless possibilities for research, education, and discovery.
Please, what image analysis software can be used for sem image analysing?special on angle calculate
Thank you for your comment. We have posted this article for general interest to scientists and are unable to recommend image analysis software. Thank you for your understanding and best wishes.
Please, what image analysis software can be used for dynamic particle counting and analysis.
Thank you for your question. Unfortunately we do not know the answer to your question, as this blog post was intended for general interest only.
i have the same question, if anyone has an answer please also inform me at ferrousllp@gmail.com
RE Neuronstudio – Automatic Neuron Tracing Super-Tool – the link is down!
Thanks for posting this information. What software would you use to control the camera and save the image?
is there a software to increase the impact of images or which can predict measure the level of impact an image can put on the viewer.
Which program would you recommend to use in undergraduate teaching labs? It needs to be super user-friendly for the students to use.
I would like my students to take their fluorescent images (captured by a smartphone…so likely a jpeg or something) and overly them to look for colocalization. Any advice would be very appreciated. Thanks!
Hi can someone help me out, to understand basic to advance in medical imagedata analysis to find out area and volume for CT SCAN image of brain
Hi Nitesh,
Unfortunately we cannot give advice for diagnostic medical procedures, as all of our products are for research use only.
Best of luck,
Patricia
hi
how can I get these software
Hi, here are the download links for
ImageJ: https://imagej.nih.gov/ij/download.html
Cell Profiler: https://cellprofiler.org/releases/
NeuronStudio: http://research.mssm.edu/cnic/tools-ns.html
VIAS: http://research.mssm.edu/cnic/tools-vias.html
L-measure: http://cng.gmu.edu:8080/Lm/
VIAS link is no longer valid. Do you know the new correct site of VIAS ?
Thanks.
I’m so sorry I can’t find a new link to VIAS anywhere!
Hi,
How to do area calculation in an image which has three different type of cells, I need to count no. of cells in each type and calculate average area in each type of cell.
I have more than 1000 images.
Please email me the link for the 5 image analysis software. Thanks,
please guide me for digital image analysis software because I am working on hyperpigmentetion
Cannot setup image j? Need help!
I am very need of software to image analysis because I work in cytogenetics.
thanks
I have a amscope camera attached to an Olympus microscope. Is there a way to work with live microscope images using imagej with this setup?
Thanks
Hi Floyd,
As we are not responsible for creating any of this software, I can only speak from my experience with these programs. I have used Micro-manager (https://micro-manager.org/) in the past on an Olympus microscope (not sure what the camera was) to acquire images. It was super simple to use. I would recommend you checkout the list of acquisition plugins on the ImageJ website (https://imagej.nih.gov/ij/plugins/#acq). You may be able to find something that meets your needs.